In Published Study, Chen Lab Elevates Understanding of Genetic Expression
ALBANY, N.Y. (April 28, 2022) — Riboswitches, natural RNA triggers that change their structure and functions when bound to small molecules, have challenged science’s understanding of biological regulation since their discovery two decades ago.
The ability of small molecules to bind to RNA and control gene expression offers another layer of regulation and the potential for direct action by compounds in the environment. Understanding how changes in RNA structure occur could lead to important therapeutic and other discoveries.

The study “Cotranscriptional RNA strand exchange underlies the gene regulation mechanism in a purine-sensing transcriptional riboswitch,” published on March 28 in the journal Nucleic Acids Research (NAR) and co-led by Alan Chen of Chemistry and the RNA Institute, takes on this challenge.
“While some riboswitches have been well studied, we lack a general understanding of how changes in RNA structure switch genetic expression from ‘on’ to ‘off,’" writes Chen and his co-lead author, chemical/biological engineer Julius Lucks of Northwestern University.
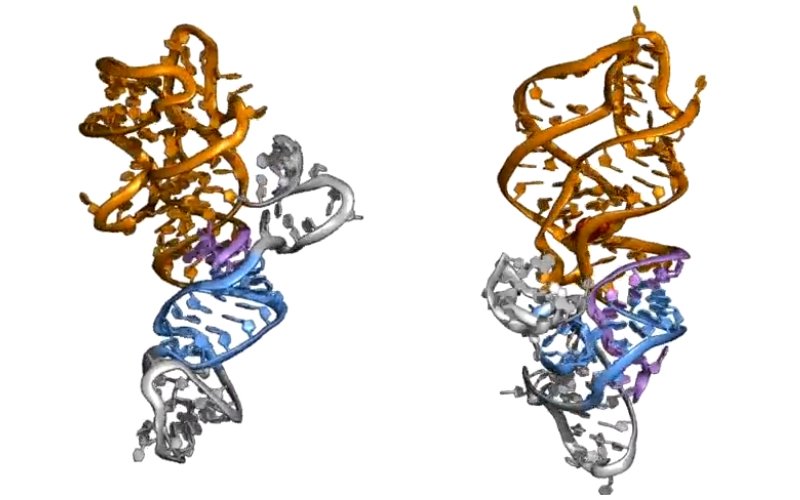
Participants on the study included two research assistants in the Chen Lab, Naomi Brandt and Elise White, who developed computational models and ran all-atom molecular dynamics simulations that confirmed one of the study's key hypotheses: that an increase in concentration of a specific metabolite — guanine — can bind specifically to the riboswitch, changing its folding pattern.
In the study, the authors propose a “strand displacement” model to explain how the RNA structure shifts between “on” and “off” states as changes occur in the concentration of small molecule ligand (a molecular group that binds to another chemical entity to form a larger complex).
“These observations help us to understand how riboswitches enable genetic decision-making,” write the authors. The data from their research provide a possible general mechanism for understanding how the competition between different strand displacement outcomes can influence RNA folding. And understanding RNA folding pathways could advance the successful design of drugs that target RNA.
“These are the first all-atom MD simulations to show the folding pathway of a purine riboswitch,” said White, who enters the PhD program in molecular biophysics at Johns Hopkins University in the fall. “They provide an elegant, mechanistic explanation for the cotranscriptional SHAPE-Seq data, and show us in molecular detail how bacteria regulate gene expression.”
Three scientists reviewing the study for NAR, a publication of Oxford University Press, nominated the manuscript for “breakthrough status,” representing the top papers NAR receives for publication, awarding it the highest possible rankings for quality, impact, and significance.
One reviewer called the study “crucial to understanding RNA function" and another said the work for the first time “addressed the gap” on how RNA folding is controlled by specific RNA sequences and how effectors influence the folding process.




